Welcome to the Hodgkin Huxley Tutorial¶
This tutorial gives an introduction to the Hodgkin-Huxley model through the use of executable example implementations in Python and NeuroML.
The aims of this tutorial are:
Provide a guide to implementing the Hodgkin-Huxley model using both Python and a NeuroML2 implementation of the same equations.
Give some background information on the electrophysiology underlying the Hodgkin-Huxley model.
The original HH tutorial was created by @joebowen on behalf of the OpenWorm project.
New interactive introduction to the HH model in a Jupyter notebook¶
A new interactive Jupyter notebook can be used to run the HH model, change the parameters of the model and display the dynamical properties of variables, without the need to write any code.
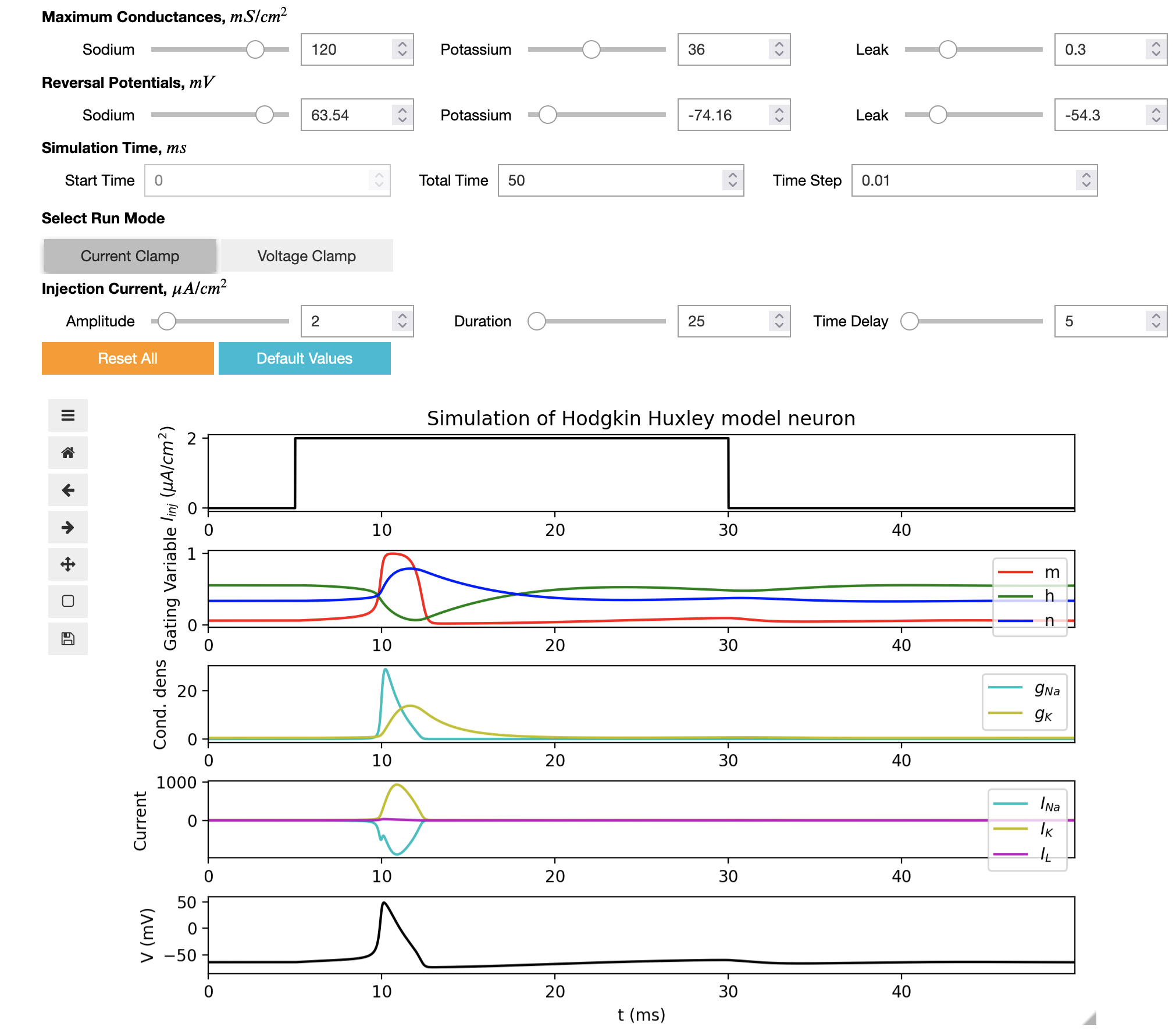
Full details can be found here with a link to execute the model in your browser. This work was carried out as part of Google Summer of Code 2022 by Rahul Sonkar.
What is the Hodgkin-Huxley model?¶
From Wikipedia: The Hodgkin–Huxley model is a mathematical model that describes how action potentials in neurons are initiated and propagated.
The model describes represents the electrical properties of excitable membranes as typical electrical circuit components. For instance, the cell’s membrane is modeled as a capacitor, and voltage-dependent conductances stand in for what are now known to be voltage-gated ion channels.
For a detailed run through of the Hodgkin-Huxley model’s electronics, math and biology, take a look at the Electrophysiology page.
After you understand the electronic model there, check out the code walkthrough to see an example implementation of the Hodgkin-Huxley model in Python, using a cell modeled in NeuroML2.
You can look at the current-voltage characteristic page to get an understanding of another biological-electronic equivalence that is useful in describing ion channel and cell models.
There are also some exercises you can complete to get a feel for the model. These can be completed using either the Python or NeuroML versions.
Table of Contents:
- Implementation of HH Model in Python and NeuroML 2
- Installing the code
- Running the model implementations
- Membrane Capacitance
- Sodium (Na) Ion Channel Variables
- Potassium (K) Ion Channel Variables
- Passive Leak Channel Variables
- Time of Simulation
- Input Current / Input Current Density
- Channel Gating Kinetics for Sodium (Na) Channel m
- Channel Gating Kinetics for Sodium (Na) Channel h
- Channel Gating Kinetics for Potassium (K) channel n
- Initial Values
- Plots
- Output of simulations
- Biological/Electronic Equivalence
- Current-voltage characteristic
- Exercises
- Hodgkin Huxley Sources